Interactive Embedded Plots with Plotly and ggplot2
Largley lifted from this r-bloggers post
install.packages("devtools") # so we can install from GitHub
devtools::install_github("ropensci/plotly") # plotly is part of rOpenScilibrary(plotly)
py <- plotly(username="jflournoy", key="mg34ox914h") # open plotly connection
# I'll change my key after this, but you can still use: plotly(username="r_user_guide", key="mw5isa4yqp")
# Or just sign up for your own account!
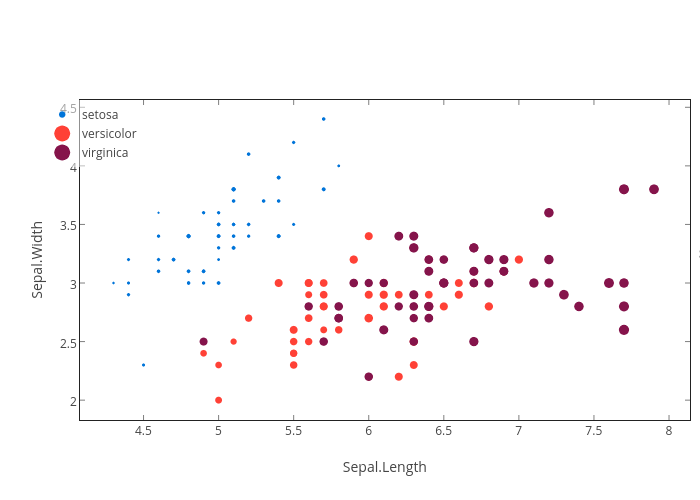
gg <- ggplot(iris) +
geom_point(aes(Sepal.Length, Sepal.Width,color=Species,size=Petal.Length))
gg#This looks a little object-oriented like python
py$ggplotly(gg)You can embed code like this (which you get from the plotly ‘share’ dialogue):
<div>
<a href="https://plot.ly/~jflournoy/16/" target="_blank" title="Sepal.Width vs Sepal.Length" style="display: block; text-align: center;"><img src="https://plot.ly/~jflournoy/16.png" alt="Sepal.Width vs Sepal.Length" style="max-width: 100%;width: 797px;" width="797" onerror="this.onerror=null;this.src='https://plot.ly/404.png';" /></a>
<script data-plotly="jflournoy:16" src="https://plot.ly/embed.js" async></script>
</div>You can also directly embed a plotly plot using a code chunk if you set plotly=TRUE for the chunk, and include session="knitr" in the call.
#Set `plotly=TRUE`
py$ggplotly(gg, session="knitr")There’s a wide world of plotly fun just waiting out there.